OBO Foundry Newsletter Issue 10
This issue reports recent decisions of the OBO Foundry Operations Committee, updates from the Editorial and Technical Working Groups, and the current status of ontology submissions under review. It also highlights infrastructure updates and ongoing discussions relevant to the broader OBO community.
The OBO Foundry continues to rely on active community engagement. Contributions through ontology development, issue tracker discussions, working group participation, tool building, and newsletter submissions remain essential to sustaining shared standards and collaborative progress.
OBO Foundry Operations Committee.
Updates
OBO Dashboard updated
The OBO Dashboard was updated (2026-03-18); ontology maintainers should review current compliance status. Ontologies that satisfy the required criteria are listed on the OBO Foundry homepage (https://obofoundry.org). Ontology developers are encouraged to review the dashboard regularly to identify areas for improvement and ensure continued compliance with Foundry standards.
The most recent dashboard status can be accessed via the OBO Dashboard page on the OBO Foundry website. (last updated 2026-03-18) here: OBO Dashboard (2026-03-18)
Decisions Made and Working Group Updates
During the Issue 10 reporting period, the Editorial Working Group and Technical Working Group finalized several documentation updates, policy refinements, and workflow changes. These actions strengthen OBO’s standardization guidance, clarify resolvability requirements, improve submission quality control, and formalize tool lifecycle management.
-
Contents of Standardization Guidelines document finalized: The Standardization Guidelines is a document developed to contain suggestions and recommendations deemed too detailed to be included as part of a Principles page. The guidelines currently include (1) Technical Considerations (addressing logical consistency after ontology reasoning, how to indicate ontology root terms, and use of standard synonym types); (2) Content Considerations (addressing preferred sources for imported terms, when to (not) import whole ontologies, and how to use NCIt terms); (3) Release Considerations (addressing the standards for different release types); and (4) Communication Considerations (addressing links to term issues). This document is not yet linked from the OBO Foundry web site, but a pre-release version is available in the Standardization Guidelines draft.
-
Principle updates clarifying IRIs and resolvability requirements: Reference to “imported from” was removed from Principle 1. Previously, “imported from” was used to indicate the ontology serving as the direct source of a term import, which could be either the primary (term-maintaining) ontology or an intermediary source. While its use is not prohibited, existing guidance recommending import from the primary source reduces its necessity.
Principle 3 was strengthened to require that the official ontology PURL, after redirection, MUST resolve to the ontology OWL file, and that PURLs for individual terms MUST resolve to dedicated, term-centric pages. These updates reinforce consistent identifier practices and improve reliability of ontology access across the OBO Foundry.
-
Terminology standardization across documentation: Mentions of “OBO Library” have been replaced with “OBO Foundry” across core documentation pages, including the Principles summary page, Principle 3, Principle 12, and the ID policy page. This update standardizes terminology across official guidance.
Ongoing Discussions
- Practice for referring to taxa not in NCBI Taxon (#434): Discussion continues regarding best practices for representing taxa not present in NCBI Taxon. The issue remains open pending further clarification.
AI-related activities in the OBO community
Artificial intelligence is under active discussion within OBO as a potential support mechanism for ontology development and curation workflows.
-
OBO Academy sessions on AI-assisted biocuration: In parallel, community-led activities are advancing the application of AI in biocuration and ontology engineering. OBO Academy sessions led by Chris Tabone demonstrated the use of large language models (Claude Code) for ontology development workflows (Part 1: https://youtu.be/pUoelp2qbDI; Part 2: https://youtu.be/UOjzTGnG_XM). These sessions illustrated practical applications, including automated term generation, ontology editing support, and workflow acceleration, while also highlighting challenges related to validation, reproducibility, and integration with existing curation standards.
-
AI in biocuration (ISB Virtual Biocuration Conference 2026): A workshop on AI in biocuration (February 9, 2026) demonstrated the use of large language models in curation workflows and knowledge representation (recording: https://youtu.be/jXHmHHLp9q0; details: https://www.biocuration.org/events/biocuration2026-virtual/). The workshop highlighted practical applications for AI-assisted curation, along with challenges related to validation, reproducibility, and integration with established ontology development standards.
-
LinkML community discussion on AI governance: A LinkML community call (February 19, 2026) addressed governance considerations for AI use, including discussion of an “AI covenant” and its potential role in defining responsible use of AI in ontology development, documentation, and data integration (recording: https://youtu.be/EOYypShdKdw).
-
AI4Curation guidance resources: Community-developed documentation provides guidance for applying AI tools in biocuration and ontology workflows (https://ai4curation.io/aidocs/), supporting more consistent and transparent adoption across projects.
-
OBO Slack channels for AI discussions: Two dedicated Slack channels (#ethical-ai, #ai) have been established to coordinate discussion of ethical, technical, and governance considerations related to AI use, indicating increasing community engagement around responsible AI integration.
Ontologies
New Ontologies Accepted in the OBO Foundry Ontology Library
Physiologically Based Pharmacokinetics Ontology (PBPKO)
- GitHub Issue: #2563
- Key Discussion: PBPKO provides structured terminology to support representation of physiologically based pharmacokinetic models and related biological processes. Following review and dashboard validation, the ontology was accepted for inclusion in the OBO Foundry. The submitter has been instructed to complete the required metadata file prior to final issue closure.
- See also: Newsletters 8-9
New ontologies currently under review
Intrinsically Disordered Proteins Ontology
- Status: Under review
- Join the discussion: GitHub Issue: #2770
- Key Discussion: Review has been completed with requested changes; the ontology is awaiting implementation of reviewer feedback before further evaluation.
Precision Fermentation Ontology
- Status: Under review
- Join the discussion: GitHub Issue: #2821
- Key Discussion: PREFER has passed NOR dashboard validation and is undergoing editorial assessment. Reviewer follow-up is pending.
- Status: Pre-registration incomplete
- Join the discussion: GitHub Issue: #2835
- Key Discussion: Required items in the pre-registration checklist remain incomplete. The submitter has been notified, and review will continue once outstanding requirements are addressed.
Clinical Metadata Exploration Ontology
- Status: Technical corrections required
- Join the discussion: GitHub Issue: #2822
- Key Discussion: Review identified invalid URIs and missing definitions. The submitter has been notified and revisions are required before further consideration.
Spotlight on well-established OBO ontologies
In this issue, we highlight The Ontology of Adverse Events (OAE) from the OBO Foundry family.
The Ontology of Adverse Events (OAE)
The Ontology of Adverse Events (OAE): A Decade of Expansion and a New Milestone.
Launched in 2011 as an OBO Foundry community ontology, the Ontology of Adverse Events (OAE) remains a critical resource for standardizing and integrating data on adverse events (AEs) following medical interventions. A recent publication in Scientific Data (January 2026; PMCID: PMC12917244) documents the ontology’s remarkable evolution over the past decade, highlighting its enhanced role in enabling computational pharmacovigilance and supporting the advancement of precision medicine. Since its last major update in 2014, OAE has undergone a substantial expansion to improve its clinical granularity and domain coverage. The ontology now contains 10,829 formally defined terms — a 251% increase from the 3,086 terms previously reported. The OAE framework now explicitly models critical determinants influencing clinical outcomes, such as administration routes, dosage parameters, and patient demographics (e.g., age), facilitating more nuanced analyses. By mining data from major sources like VAERS and FAERS and integrating with regulatory vocabularies like MedDRA, OAE effectively bridges the gap between raw clinical reports and structured computational analysis. To broaden its utility within the global scientific community, developers have also introduced “Chinese Translation” annotations for new terms, supporting OAE’s applications in monitoring post-immunization adverse events in China. By providing a logically structured and interoperable platform for adverse event representation, OAE continues to empower cross-disciplinary analysis of adverse event patterns, etiological factors, and outcome trajectories, ultimately fostering safer medical interventions and advancing translational research.
Updates on well-established OBO ontologies
MONDO (Mondo Disease Ontology)
The Mondo Disease Ontology is now available within Epic Electronic Health Record (EHR) systems through integration with IMO Health’s terminology platform. As part of Epic’s February 2026 software update, health systems using IMO Core can document diagnoses using Mondo-linked terms directly in clinical workflows.
IMO Health has mapped nearly 5,000 new rare disease diagnoses and aligned more than 25,000 existing clinical terms to standardized Mondo identifiers. This integration enables clinicians to continue using familiar terminology while ensuring that documentation is connected to unified, computable Mondo IDs behind the scenes.
The expansion significantly improves rare disease representation in clinical systems. Coverage of rare genetic disease diagnoses increases from 51% to 75% compared to ICD-10 alone, while maintaining greater than 99% precision. Research conducted at Cincinnati Children’s further demonstrated that enhanced terminology support identified 5.5 times more rare diseases and 20% more patients than standard ICD-10 coding alone.
Embedding Mondo into routine clinical documentation strengthens the alignment between research-grade disease definitions and point-of-care workflows. Structured capture of rare disease data supports improved cohort identification, clinical trial matching, longitudinal analysis, and downstream analytics, including artificial intelligence applications that depend on standardized, high-quality clinical data.
Members and Volunteers
The OBO Foundry is honored to highlight valued community members in each issue. This time, we feature Valerie Wood and Shawn ZK Tan.
Valerie Wood

Valerie Wood is Project Manager of PomBase (www.pombase.org), the comprehensive knowledgebase for Schizosaccharomyces pombe, a widely used model eukaryote. She has over 30 years of experience in fission yeast biocuration, beginning with genome sequence feature annotation and subsequently expanding into ontology-driven curation of molecular functions, genetic and physical interactions, phenotypes, and, more recently, GO-CAM pathway models.
Val has been a major contributor to the development of the Gene Ontology since its inception. She also maintains the Fission Yeast Phenotype Ontology (FYPO), originally developed by Midori Harris, and contributes regularly to several community ontologies, including Mondo Disease Ontology, the Sequence Ontology, and ChEBI (Chemical Entities of Biological Interest).
She recently published a perspective article titled “Empowering Biological Knowledgebases: Advances in Human-in-the-Loop AI-Driven Literature Curation” (DOI: 10.1093/bioadv/vbag028), with colleagues from the ELIXIR Biocuration Focus Group. This work reviews how artificial intelligence (particularly large language models) is being applied to support literature curation, outlines key challenges, and provides practical recommendations for future development.
PomBase is actively integrating AI into its biocuration workflows, including the use of ai4curation tools developed by Chris Mungall and colleagues to accelerate ontology development, as well as exploring AI-based approaches for extracting ontology-driven annotations from the literature.
Shawn ZK Tan

Shawn is the citizen technology lead in GSK Asia Research & Development and works as an independent consultant operating under his own consulting practice, SignaMind. Shawn has also been part of the OBO Foundry Operations Committee since 2022. Starting his career as a lab scientist, Shawn only started work on biomedical ontologies after his PhD when he joined EMBL-EBI under David Osumi-Sutherland as an ontology editor, developing an ontology to support the data coming out from single cell mapping of the brain. During his time at EBI, Shawn actively contributed to the Cell Ontology (CL), Uberon, the Phenotype And Traits Ontology (PATO), and the Relation Ontology (RO). After his time at EMBL-EBI, Shawn joined Novo Nordisk first as a Senior Research Data Steward embedded in both research projects and ontology development. He subsequently took up a role as a Business Capability Lead where he drove the development and operationalisation of the ontology strategy. During his time at Novo Nordisk, Shawn was a strong advocate for using OBO Foundry ontologies and tools in industry, and for industry taking a bigger role in supporting the OBO community, embedding tools into pipeline and writing about the mutual benefits that exist in such a relationship. In his current roles, he continues advocating for the use of open standards and tools in industry and aims to be a strong advocate for Asia’s data for global use, with ambitions to build up communities and representation in Asia for such purposes. After experiencing the challenges of fragmented data across modern organizations and how they tend not to get the most out of using bio-ontologies, Shawn also started a consultancy, SignaMind, which focuses on helping organizations build up their semantic capabilities, and doing so with strong business value in mind. With SignaMind, he is currently working with The Novo Nordisk Foundation Biotechnology Research Institute for the Green Transition (BRIGHT) to build up their semantic capabilities, starting with an ontology for precision fermentation (PREFER).
Spotlight on Research and practice in the OBO community
“Curate This!” Podcast Series Launched by ISB
International Society for Biocuration (ISB) has launched a new podcast series titled “Curate This!”, hosted by Susan Bello and featuring contributions from Charlie Hoyt.
The series highlights perspectives from the biocuration community, exploring the people, practices, and evolving challenges that shape knowledge representation in the life sciences. Topics include ontology development, curation workflows, interoperability, and community infrastructure.
The podcast provides an accessible platform for discussing both practical and conceptual aspects of biocuration, with relevance for ontology developers, knowledge engineers, and data integration specialists across the OBO ecosystem.
The inaugural episode is available on YouTube
The DO-KB knowledgebase 2026 update: expanding programmatic and language access
This recent article reports the 2026 update of the Human Disease Ontology Knowledgebase (DO-KB) in Nucleic Acids Research (Database Issue). Baron et al. describe substantial expansion in disease representation together with major upgrades to programmatic access and multilingual delivery (published online 26 November 2025; Issue 6 January 2026). The update strengthens interoperability through an OpenAPI (OAS 3.1) compliant API, an expanded and reorganized SPARQL Sandbox (now 55 queries with pagination and federated examples), and enhanced cross-resource mappings using structured match designations and SSSOM outputs.
A central contribution is internationalization: the Disease Ontology implemented language tags across labels, definitions, and synonyms and released Spanish-language ontology artifacts alongside bilingual website navigation. This work provides a practical model for multilingual delivery within the OBO ecosystem. The article also documents governance-through-practice improvements aligned with OBO operational priorities, including a clinician-led nosology education program that feeds curated updates back into the ontology and infrastructure modernization (e.g., Neo4j upgrade) to improve performance, sustainability, and long-term accessibility.
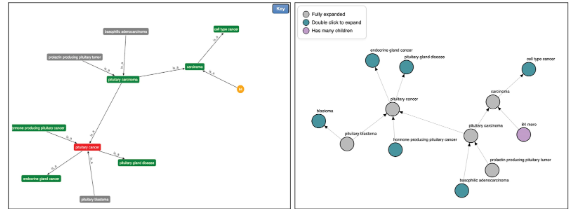
Figure from Baron JA, Sánchez-Beato Johnson CM, Schor MA, Olley D, Nickel L, Felix V, Bello SM, Greene C, Lichenstein R, Bisordi K, et al. The DO-KB knowledgebase 2026 update: expanding programmatic and language access. Nucleic Acids Research. 2026;54(D1):D1425–D1436. https://doi.org/10.1093/nar/gkaf1213
Epilepsy disease classification: a community effort to enhance the Mondo Disease Ontology
This recent article reports a community-driven effort to improve the representation and classification of epilepsy within the Mondo Disease Ontology. Vasilevsky et al. describe a series of expert-led workshops that aligned Mondo’s epilepsy hierarchy with the International League Against Epilepsy (ILAE) classification, resulting in substantial revisions to disease structure, terminology, and classification logic (published 16 February 2026). Updates include the addition of new epilepsy types, reclassification of syndromes and etiologies, refinement of hierarchical relationships, and extensive updates to synonyms, definitions, and cross-references to improve clinical relevance and interoperability.
A central contribution is the integration of clinical and ontology expertise to produce a more granular and standardized framework for epilepsy representation. The revised structure supports flexible annotation across levels of diagnostic specificity and improves data integration across resources. The work also highlights ongoing challenges, including defining epilepsy syndromes, distinguishing disease from phenotype, and modeling genetic and temporal aspects of disease. These updates strengthen Mondo’s role in supporting rare disease research, clinical decision-making, and data harmonization across platforms such as RDCA-DAP and ClinGen, while establishing a foundation for continued community-driven refinement.
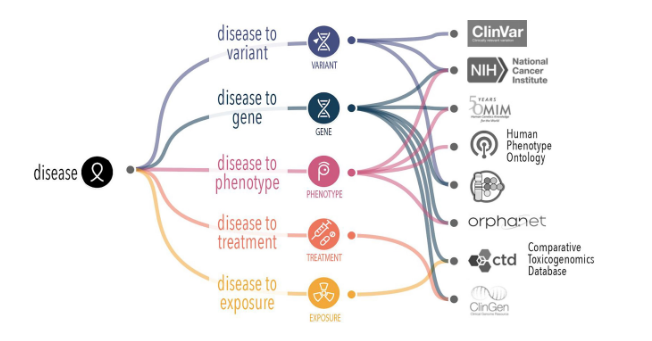
Figure from Vasilevsky N, Gehrke S, Mullen K, Barua S, Braun I, Brünger T, Coughlin C II, Ivaniuk A, Korn D, Lal D, Marsh S, O’Loughlin E, Olson D, Shwetar Y, Sofocleous C, Vogel-Farley V, Grabenstatter H, Haendel M, Mungall C, Toro S. Epilepsy disease classification: a community effort to enhance the Mondo Disease Ontology. Database. 2026;2026:baag004. https://doi.org/10.1093/database/baag004
ChEBI: re-engineered for a sustainable future
This recent article, “ChEBI: re-engineered for a sustainable future,” describes a full modernization of the Chemical Entities of Biological Interest (ChEBI) database and ontology, motivated by the growing maintenance burden of its legacy infrastructure. Malik et al. report a complete overhaul of ChEBI’s platform database, website, web services, and submission tooling—to improve sustainability, scalability, and long-term operability for a global user base (published online 28 November 2025; Issue 6 January 2026). Key infrastructure changes include migrating from Oracle to PostgreSQL, containerized deployment via Kubernetes, and redesigned web interfaces built for responsive use and improved discovery.
The update also improves programmatic and user-facing access to ChEBI content by replacing legacy SOAP services with modern REST APIs (with interactive Swagger UI documentation), upgrading text search from Lucene to Elasticsearch, and updating chemical structure search to RDKit-based tooling (including dedicated support for substructure and similarity workflows). For ontology and data quality, the release introduces standardized identifiers and relationships (including broader alignment with OBO Relation Ontology), adds new export formats such as OBO JSON Graph, and applies ROBOT-based QC checks—reducing legacy ontology errors and improving consistency of metadata and annotation properties.
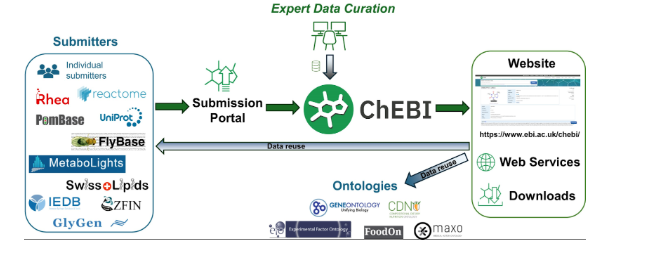
Figure from Malik A, Arsalan M, Moreno C, Mosquera J, Félix E, Kizilören T, Muthukrishnan V, Zdrazil B, Leach AR, O’Boyle NM. ChEBI: re-engineered for a sustainable future. Nucleic Acids Research. 2026;54(D1):D1768–D1778. https://doi.org/10.1093/nar/gkaf1271
Spotlight on Research: Now Published
Several research papers previously featured in the OBO Newsletter as preprints have now been published in journals.
The Ontology of Adverse Events in 2025
This recent article reports substantial updates to the Ontology of Adverse Events (OAE), an OBO Foundry ontology designed to standardize and integrate adverse events associated with medical interventions. The work focused on expanding clinical representation, improving semantic structure, and strengthening interoperability to support computational analysis across biomedical domains.
Over the past decade, OAE has expanded from 3,088 terms to more than 10,800 formally defined terms, reflecting significant growth in domain coverage and clinical granularity. New content includes detailed representation of patient anatomic regions, clinical manifestations, and pathological processes, with terms derived from sources such as VAERS, FAERS, MedDRA, and published studies. The ontology also captures key determinants influencing adverse event outcomes, including administration routes, dosage parameters, and demographic variables, enabling structured analysis of clinical and epidemiological patterns.
Interoperability has been strengthened through integration with existing ontologies and knowledge bases, including alignment with BFO as an upper-level ontology and mapping to resources such as the Human Phenotype Ontology, Uberon, and Gene Ontology. OAE supports the development of domain-specific extensions, such as drug- and disease-specific adverse event ontologies, and enables semantic integration across pharmacovigilance datasets and biomedical knowledge systems.
Infrastructure and annotation practices have also been refined. Standardized annotation schemas ensure consistent use of labels, definitions, and metadata, while tools such as OntoFox, OntoBee, and OntoRat support ontology development and reuse. Internationalization efforts include the introduction of Chinese translations and broader adoption in global research contexts, particularly in China’s biomedical data infrastructure initiatives.
Together, these developments position OAE as a comprehensive and interoperable framework for representing adverse events, supporting pharmacovigilance, precision medicine, and cross-domain data integration within the OBO ecosystem.
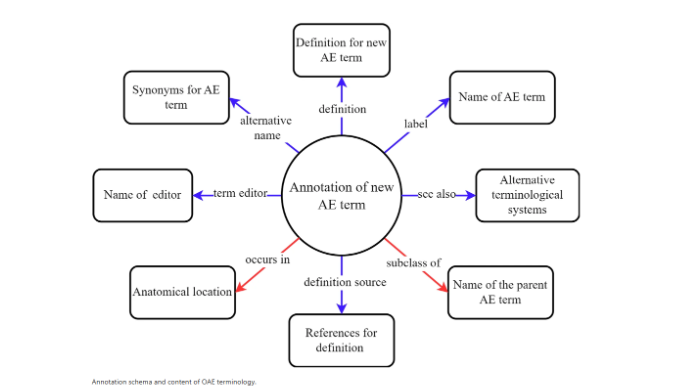
Figure from Chenchen Pan, Qiuyue Yang, Xue Zhang, Shuyue Luo, Yongqun He, Jiangan Xie. The Ontology of Adverse Events in 2025. 2026 January 13. https://doi.org/10.1038/s41597-026-06584-x
Spotlight on OBO Principles
Principle 8: Documentation
To ensure transparency, traceability, and long-term sustainability, OBO Foundry ontologies must provide clear and comprehensive documentation. Principle 8 requires ontology owners to document the full ontology life cycle and to address multiple audiences, including users and developers. Documentation serves as an indicator of ontology quality and is expected to evolve alongside the ontology, including updates and release records as the resource matures.
Documentation should cover several complementary areas. Global ontology documentation includes metadata describing the ontology as a whole, such as creators, maintainers, license, and version information. Local ontology documentation addresses representational units (classes and relations), including justification of metadata elements, explanation of axiomatization, and clarification of whether textual definitions are manually authored or generated using patterns or software assistance. User documentation should describe the ontology’s purpose, scope, intended use cases, and access methods, potentially including SPARQL query examples, training materials, tutorials, publications, and recorded presentations. Developer documentation should outline contribution workflows (e.g., via a CONTRIBUTING.md file), licensing terms, version control practices, release procedures, deprecation and conflict resolution policies, batch term submission processes, continuous integration, interaction with orthogonal resources, and the use of automated agents for quality control and maintenance.
Implementation requirement: If a term originally defined in ontology A is adopted into ontology B with a new identifier, the annotation assertion rdfs:isDefinedBy MUST be added to record provenance.
In OWL (Turtle serialization), this is expressed as:
<http://purl.obolibrary.org/obo/A_123> rdfs:isDefinedBy <http://purl.obolibrary.org/obo/b.owl> .
In OBO format, this is expressed as:
property_value: isDefinedBy http://purl.obolibrary.org/obo/b.owl
This requirement ensures traceability of adopted terms across ontologies.
Validation: Principle 8 is automatically validated through the OBO Dashboard. Documentation presence and completeness are assessed as part of overall ontology soundness evaluation.
Open considerations include clarifying minimal required metadata, defining minimal documentation thresholds, and articulating the role of peer-reviewed publications in satisfying documentation expectations.
For more details, see the OBO Foundry documentation on Principle 8.
Spotlight on Tools: Recent updates
- Retirement of
[funowl](https://github.com/Harold-Solbrig/funowl): Thefunowllibrary has been officially retired. A migration guide has been published to support users in transitioning to alternative tooling. This formal retirement clarifies maintenance expectations and helps prevent reliance on unsupported infrastructure. - Ontology Development Kit (ODK) minor 1.6.1 release with various tool updates
- ROBOT 1.9.10 release with updates to obographs format, prefix injections into OBO format files, a global –input-format option to avoid cycling through all possible parsers with the OWL API and more.
Events and News
Updates on past events
Core Ontology for Biology and Biomedicine (COB) Workshop (January 26, 2026)
The workshop reviewed progress in COB implementation across the OBO Foundry and identified key barriers to adoption. Reported challenges included uncertainty around required COB imports, lack of clear criteria for full COB alignment, and dependencies on RO relations not represented in COB. Planned actions include improving COB documentation, clarifying release artifacts for alignment, and defining alignment standards for ontologies at different stages of development. Coordination between RO and COB was identified as a priority for future work.
Upcoming events
Conferences at ISMB 2026 (Washington, DC)
Three ontology-relevant meetings will be held in July 2026:
- Bio-Ontologies and Knowledge Representation (BOKR) – July 13–14
- Bioinformatics Open Source Conference (BOSC) – July 14–15
- A joint BOKR/BOSC session will be held for half a day.
- Abstracts on open-source approaches to knowledge representation may be submitted to either track via ISMB EasyChair.
- Deadline for talk/poster abstracts: April 9 (not expected to be extended).
- In-person presentation is required unless a medical or caregiver exemption applies.
- International Conference on Biomedical Ontology (ICBO) – July 13–18
- Workshops and tutorials are expected during the first part of the week.
- Submission details are pending clarification.
Biocuration 2026 (Virtual)
- The 2026 Virtual Biocuration Conference Series brings together biocurators, database developers, data scientists, and the broader research community in an accessible and flexible format.
Workshops with Upcoming Deadlines
- ESWC 2026 (Dubrovnik, Croatia, May 10–14)
- Workshop: LLMs and Knowledge Graphs (LLM4KGOE)
- Invited speakers are being sought (not funded; registration and travel required).
- Deadline: February 24 (may be extended).
- ICHI 2026 (Minnesota, USA, June 1–2)
- Workshop: Food and Health Knowledge Graphs Using Large Language Models (LLMs)
- Short and long papers will be published in the proceedings.
- Deadline: March 15.
- Invited speakers are welcome (not funded).
OBO Academy: Seminar Series (ongoing)
The OBO Academy Seminar Series, organized by members of the Monarch Initiative and open to all, is active again after the summer break. The series offers training and discussions on ontology-related tools and practices. To join, please reach out in the OBO Slack channel #obo-academy.
Upcoming seminars:
- None planned at this time.
The full schedule and materials are available at https://oboacademy.github.io/obook/courses/monarch-obo-training/
You can also vote for or request new training topics at https://github.com/OBOAcademy/obook/discussions/categories/tutorial-requests
Ways to be part of the OBO Foundry community
There are many ways to be part of the OBO Foundry community, beyond using our website to find ontologies of interest. For example:
- Join the obo-discuss mailing list
- If you are interested in the technical aspects, you can also join the obo-tools mailing list
- Join the conversation in our Slack workspace.
- OBO Foundry RSS feed at https://obofoundry.org/feed.xml
- Use our public issue tracker to report a problem you discovered on obofoundry.org or request a new feature
- Note that most issues relating to individual ontologies (e.g., a request to add a new term) should be instead added to the issue tracker for the specific ontology
- Propose a fix to an issue you see on our issue tracker (this is done via a GitHub Pull Request, which will be checked and approved by someone in the Foundry). Since all of our code is publicly readable, you may be able to pinpoint where a bug fix needs to go.
- Request that your ontology be considered for inclusion in the Foundry
- Suggest content for our next newsletter
- If you want to take your contributions to OBO Foundry to the next level, you can nominate yourself to be considered for the OBO Operations Committee. There are many ways in which you can contribute, including assisting with the production of this newsletter.